Abstract
Aims/hypothesis
Diabetes and hyperlipidaemia are common comorbidities in people with hypertension. Despite similar protective effects on CVD, different classes of antihypertensive drugs have different effects on CVD risk factors, including diabetes, glucose metabolism and lipids. However, these pleiotropic effects have not been assessed in long-term, large randomised controlled trials, especially for East Asians.
Methods
We used Mendelian randomisation to obtain unconfounded associations of ACE inhibitors, β-blockers (BBs) and calcium channel blockers (CCBs). Specifically, we used genetic variants in drug target genes and related to systolic BP in Europeans and East Asians, and applied them to the largest available genome-wide association studies of diabetes (74,124 cases and 824,006 controls in Europeans, 77,418 cases and 356,122 controls in East Asians), blood glucose levels, HbA1c, and lipids (LDL-cholesterol, HDL-cholesterol and triacylglycerols) (approximately 0.5 million Europeans and 0.1 million East Asians). We used coronary artery disease (CAD) as a control outcome and used different genetic instruments and analysis methods as sensitivity analyses.
Results
As expected, genetically proxied ACE inhibition, BBs and CCBs were related to lower risk of CAD in both ancestries. Genetically proxied ACE inhibition was associated with a lower risk of diabetes (OR 0.85, 95% CI 0.78–0.93), and genetic proxies for BBs were associated with a higher risk of diabetes (OR 1.05, 95% CI 1.02–1.09). The estimates were similar in East Asians, and were corroborated by systematic review and meta-analyses of randomised controlled trials. In both ancestries, genetic proxies for BBs were associated with lower HDL-cholesterol and higher triacylglycerols, and genetic proxies for CCBs were associated with higher LDL-cholesterol. The estimates were robust to the use of different genetic instruments and analytical methods.
Conclusions/interpretation
Our findings suggest protective association of genetically proxied ACE inhibition with diabetes, while genetic proxies for BBs and CCBs possibly relate to an unfavourable metabolic profile. Developing a deeper understanding of the pathways underlying these diverse associations would be worthwhile, with implications for drug repositioning as well as optimal CVD prevention and treatment strategies in people with hypertension, diabetes and/or hyperlipidaemia.
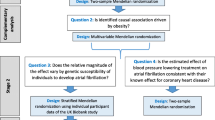
Graphical abstract

Similar content being viewed by others

Introduction
Hypertension and diabetes are well-established risk factors for CVD, a leading cause of mortality. The cardiovascular benefits of BP control, including antihypertensive drugs, are well documented [1, 2]. Despite their similar cardiovascular benefits, different classes of antihypertensive drugs have shown different off-target effects on type 2 diabetes (‘diabetes’ for short), glucose metabolism and lipid profile, which are well-established CVD risk factors. ACE inhibitors have been recommended as the first-line treatment in the treatment of hypertension in patients with diabetes due to their beneficial effects on renal function, followed by calcium channel blockers (CCBs) [3]. ACE inhibitors have also shown benefits in lowering blood glucose levels, possibly lowering the risk of diabetes in an RCT [4]. In contrast, β-blockers (BBs), a commonly used antihypertensive drug in uncomplicated hypertension, are not recommended in people with the metabolic syndrome, in whom there are concerns about unexpected off-target effects on glucose metabolism and lipid profile based on observational studies [5, 6] and small clinical trial evidence [7]. However, the observed associations are inconsistent [8, 9], and may be open to residual confounding by socioeconomic position and health status. Evidence from RCTs is not clearly established due to the relatively short follow-up in trials with diabetes as the primary outcome (such as for ACE inhibitors) [4] and ethical considerations precluding examination of expected harms as a primary outcome in large RCTs (such as for BBs). In particular, the evidence regarding the metabolic effect of CCBs is quite limited. In an interventional study, there was an increase in LDL-cholesterol after taking CCBs [10], which has not been assessed in large RCTs comparing CCBs with placebo.
Current RCTs have mainly been conducted in people of European ancestry, but the metabolism of antihypertensive drugs may be different in East Asian participants, and off-target effects may also differ by population. For example, Chinese participants were more likely to have cough, a common side effect of ACE inhibitors, than Europeans [11]. Although diabetes is more prevalent in East Asians than in Western populations [12], evidence from East Asians is lacking.
Mendelian randomisation (MR) studies, which can foreshadow the results of RCTs, provide an approach to assess long-term off-target effects [13]. As genetic variants are randomly allocated at conception and not affected by socioeconomic position or other confounders, the study design minimises residual confounding. In contrast to conventional MR for an exposure, the instrument proxying drug effects in MR can be constructed based on the drug target genes, rather than genetic variants from across the genome [14]. Using genetic variants in the drug target genes as proxies [15,16,17,18], we examined the role of ACE inhibitors, BBs and CCBs in diabetes and metabolic traits in people of both European and East Asian ancestries based on the large genome-wide association studies (GWAS).
Methods
Study design
By means of an MR study in Europeans and East Asians, we assessed the role of ACE inhibitors, BBs and CCBs in diabetes, glycaemic traits, and the levels of LDL-cholesterol, HDL-cholesterol and triacylglycerols. For glycaemic traits, we evaluated blood glucose levels and HbA1c, as they are commonly used in diabetes diagnosis. We additionally assessed their roles in insulin levels without adjustment for BMI in Europeans, and the role in BMI in both ancestries given it is a potential mediator between antihypertensive drugs and diabetes [19]. In European participants, we applied the genetic proxies for ACE inhibitors, BBs and CCBs to large cohorts with extensive genotyping. In East Asian participants, we used genetic proxies identified from Biobank Japan, and applied them to the large GWAS of diabetes, glucose, HbA1c and lipids. For comprehensiveness, we further assessed their roles in lipid-related variables including lipid fractions, 14 lipoprotein subclasses and apolipoproteins in European participants. Coronary artery disease (CAD) was used as a control outcome to assess the validity of the genetic instruments. For comparison with evidence from RCTs, we also conducted a systematic review and meta-analysis of RCTs on the use of ACE inhibitors, BBs and CCBs in new-onset diabetes.
Exposure
For the genetic instruments in European participants, we used genetic proxies for ACE inhibitors, BBs and CCBs provided by previous studies [15,16,17,18]. Specifically, previous studies identified the genes encoding pharmacological targets for these antihypertensive drugs in DrugBank (ACE for ACE inhibitors, ADRB1 for BBs, and CACNA1D, CACNA1F, CACNA2D1, CACNA2D2, CACNA1S, CACNB1, CACNB2, CACNB3, CACNB4, CACNG1 and CACNA1C for CCBs) [16, 17]. Genetic variants that are present in these genes or regulatory gene regions, and also associated with systolic BP (SBP) in GWAS meta-analysis of UK Biobank and data from the International Consortium of Blood Pressure, were selected as proxies for the antihypertensive drug. The GWAS meta-analysis was performed in up to 757,601 participants, with adjustment for age, age squared, sex, BMI, principal components and study-specific covariates [16, 17]. Given that the associations were adjusted for heritable covariates, such as BMI [20], we also used another set of genetic instruments that are located near (± 200 kb) or within these genes and related to SBP based on the GWAS in the UK Biobank (Pan UKB) [15], with no adjustment for BMI. As previously [18], we used genetic variants with r2 <0.01 for both sets of genetic proxies, with reference to the European 1000 Genomes panel (http://fileserve.mrcieu.ac.uk/ld/1kg.v3.tgz). Details of data sources are shown in ESM Table 1. The genetic proxies are shown in ESM Table 2. We also checked the relevance of these genetic proxies to the use of corresponding antihypertensive drug, using summary statistics in the UK Biobank. For ease of comparison, for both sets of genetic proxies for antihypertensive drugs, we used the genetic associations with SBP obtained from the UK Biobank summary statistics. The strength of each genetic variant was assessed using the F-statistic, obtained using an established approximation based on the genetic association with SBP and the SE; only genetic variants with F-statistic >10 were used (ESM Table 2).
To assess the associations in East Asians, we used genetic variants that are located near (± 200 kb) or within these drug target genes, and also associated with SBP in a GWAS in Biobank Japan, as instruments. Biobank Japan is a hospital-based registry that collected DNA, serum and clinical information from approximately 200,000 patients, of East Asian descent from 2003–2007 [21]. The GWAS of SBP in 136,597 participants controlled for sex, age, age squared, the top ten principal components, status of 47 diseases and smoking status [21]. We selected genetic variants with F-statistic >10 and with a linkage disequilibrium (LD) threshold of r2 < 0.1 based on the East Asian 1000 Genomes panel. As the genetic instruments were selected based on a smaller GWAS of SBP (~0.1 million) than in Europeans (>0.7 million), we used a less stringent LD threshold than for Europeans to increase the comparability to Europeans. The details of the genetic proxies are shown in ESM Table 3. We also obtained their associations with SBP in Chinese participants (n ≤ 11,816), controlling for age, age squared, gender and BMI [22].
Antihypertensive drugs have a common effect of lowering BP. To assess the metabolic effect of SBP for comparison with that of antihypertensive drugs, we also assessed the role of genetically predicted SBP in Europeans and East Asians. In contrast to the investigation of genetically proxied antihypertensive drugs that used genetic variants in specific drug target genes, the investigation of genetically predicted SBP used genetic variants that were related to BP at genome-wide significance across the whole genome [14, 16]. A more stringent LD threshold was usually used in instrument selection for SBP than for antihypertensive drugs [14, 16]. Specifically, genetic proxies for SBP were obtained from the published GWAS of SBP in UK Biobank for Europeans, as described above [15], and from a large published GWAS in East Asians [23], with an LD threshold of r2 of 0.001 with reference to the European and East Asian 1000 Genomes panels, respectively.
Outcomes
In Europeans, genetic associations with diabetes were obtained from the large GWAS of type 2 diabetes in DIAGRAM, which includes 898,130 participants of European ancestry (74,124 cases and 824,006 controls) [24]. In East Asians, genetic associations with diabetes were obtained from a GWAS meta-analysis in 433,540 East Asian individuals (77,418 cases and 356,122 controls) [25].
We also obtained genetic associations with glycaemic traits and lipids from the UK Biobank and large consortia. UK Biobank is a large, ongoing, prospective cohort study, currently with a median follow-up of 11.1 years [26]. It recruited 502,713 people (aged 40–69 years, mean age 56.5 years, 45.6% men) from 2006–2010 in England, Scotland and Wales, 94% of whom had self-reported European ancestry. Additionally, genetic associations with glucose and HbA1c were obtained from MAGIC (140,595 for fasting glucose and ≤145,579 for HbA1c). Genetic associations with insulin and BMI were obtained from MAGIC and GIANT, respectively (ESM Table 1). Genetic associations with lipids were obtained from both UK Biobank and the Global Lipids Genetics Consortium (188,577 participants of European descent and 7898 of non-European descent [27]). Genetic associations with lipid fractions, 14 lipoprotein subclasses and apolipoproteins were further assessed by metabolomics analyses performed in a randomly selected subset of UK Biobank [28].
In East Asians, genetic association with glucose, LDL-cholesterol, HDL-cholesterol and triacylglycerols was obtained from Biobank Japan in 93,146 people, with adjustment for sex, age, the top ten principal components and status of 47 diseases [21]. Genetic association with HbA1c was obtained from both Biobank and MAGIC (ESM Table 1). Genetic association with BMI was obtained from Biobank Japan in 158,284 participants. We also obtained genetic associations with glucose (n = 6063) and lipids (n ≤ 10,341) from Chinese participants, with adjustment for age, age squared and study-specific covariates [29].
We used CAD as a control outcome, because antihypertensive drugs are known to have cardiovascular protective effect. In Europeans, the genetic association with CAD was obtained from a large genome-wide association study in 34,541 CAD cases and 261,984 controls from UK Biobank, followed by replication in 88,192 cases and 162,544 controls from CARDIoGRAMplusC4D [30]. In East Asians, genetic association with CAD was obtained from Biobank Japan and Chinese cohorts. Specifically, we used genetic associations with CAD (myocardial infarction, stable angina and unstable angina) in 204,745 Japanese participants (cases = 21,611, controls = 183,134) [31] and 11,389 Chinese participants (3570 cases and 7819 controls) [32].
Statistical analysis
MR estimates were based on SNP-specific Wald estimates, i.e. the genetic association with the outcome divided by the genetic association with the genetic proxies for antihypertensive drugs (proxied by the genetic association with SBP, in units of 1 mmHg). We meta-analysed the Wald estimates for each genetic variant using inverse variance weighting with multiplicative random effects when multiple genetic variants were used. We assessed the role of SBP using the same method. To maximise power, the MR estimates for glucose from MAGIC and UK Biobank in European participants were meta-analysed together as the main analysis. The MR estimates for lipids from the Global Lipids Genetics Consortium and UK Biobank were meta-analysed. In East Asians, the MR estimates for glucose and lipids from Japanese and Chinese participants were meta-analysed. We used Bonferroni correction to account for the multiple testing, with the p value cut-off at 0.05/9 (six primary outcomes, one control outcome and two secondary outcomes)/3 (three antihypertensive drugs) = 0.002. Power calculation was performed in Europeans and East Asians. Power calculations were based on the approximation that the sample size of an MR study is the sample size for exposure on outcome divided by the r2 for genetic proxies on exposure [33, 34].
To assess the validity of these genetic instruments, we evaluated their association with CAD; an inverse association with CAD was expected. We also used MR Steiger, which enables inference of the causal direction, by calculating and comparing the variance explained by the genetic instrument in each outcome and in SBP [35]. In sensitivity analysis, we also used another set of genetic proxies in Europeans derived from the UK Biobank (ESM Table 2). In addition, we used various methods to control for pleiotropy, including a weighted median [36] and Mendelian Randomisation Pleiotropy RESidual Sum and Outlier (MR-PRESSO) [37], where applicable. The weighted median estimate is robust to invalid instruments and able to provide consistent estimation even when up to 50% of the weight is from invalid SNPs [36]. MR-PRESSO identifies outliers with potential horizontal pleiotropy and provides a corrected estimate after removing these outliers [37]. All statistical analyses were performed using R version 4.0.1 (R Foundation for Statistical Computing, Austria), and the R package ‘MendelianRandomization’.
Systematic review and meta-analysis
This systematic review and meta-analysis was registered in Research Registry (number: reviewregistry1239), providing details of the literature search, selection criteria, data extraction and data analysis. Briefly, we searched in PubMed for RCTs reporting the effect of ACE inhibitors, BBs or CCBs on new-onset diabetes, using (‘angiotensin-converting enzyme inhibitors’ or ‘ACE inhibitors’ or ‘beta-adrenergic blockers’ or ‘beta-blockers’ or ‘calcium channel blockers’) and ‘hypertension’ and ‘diabetes’ and ‘trial’ in any field. We also checked the references of relevant systematic reviews and meta-analyses. We included RCTs comparing ACE inhibitors vs CCBs or placebo, comparing BBs vs CCBs or placebo, and comparing CCBs vs placebo, with follow-up of no less than one year. The random-effects model in ‘metafor’ package in R was used for the meta-analysis. We used funnel plots and ‘trim and fill’ to assess publication bias, i.e. missing studies, utilising the ‘funnel’ and ‘trimfill’ functions from the ‘metafor’ package. We assessed the quality using an established tool [38], and excluded low-quality studies where applicable.
Results
Genetic proxies for antihypertensive drugs on diabetes and blood glucose levels
The genetic proxies for Europeans and East Asians are shown in ESM Tables 2 and 3. As expected, the genetic proxies for ACE inhibitors, BBs and CCBs had an inverse association with CAD in both Europeans and East Asians (ESM Fig. 1), which supported the validity of the genetic instruments. Most genetic proxies were also related to the use of the corresponding antihypertensive drug (ESM Table 2). Using these genetic instruments, genetically predicted ACE inhibition was associated with a lower risk of diabetes in Europeans (OR 0.85 per mmHg lower SBP, 95% CI 0.78–0.93) and East Asians (OR 0.94, 95% CI 0.90–0.99) (Fig. 1). Similar to the association for diabetes, there was an inverse association of genetically predicted ACE inhibition with blood glucose levels in meta-analysis of the UK Biobank and MAGIC (Fig. 1), with consistent associations in the UK Biobank and MAGIC (ESM Fig. 2). The association in East Asians was in the same direction, but included the null (Fig. 1), possibly due to the relatively small sample size in East Asians (~0.1 million) compared with Europeans (~0.5 million). Genetically predicted ACE inhibition was also related to lower HbA1c (Fig. 1) and BMI (ESM Table 4) in Europeans but had null associations in East Asians (Fig. 1 and ESM Table 4). In contrast, genetic proxies for BBs were associated with a higher risk of diabetes (OR 1.05, 95% CI 1.02–1.09) in both Europeans and East Asians (OR 1.03, 95% CI 1.00–1.05). Genetic proxies for BBs were also related to higher HbA1c in Europeans (Fig. 1). For CCBs, no association with diabetes or blood glucose levels in Europeans was observed, although the association in East Asians showed a nominal significance. All three drugs had null association with insulin in Europeans (ESM Table 4).
Genetic proxies for antihypertensive drugs on lipids
As shown in Fig. 2, genetically predicted ACE inhibition was associated with higher HDL-cholesterol using meta-analysis of UK Biobank and the Global Lipids Genetics Consortium in Europeans. There was a null association in East Asians, although the direction of association was consistent. In both Europeans and East Asians, genetic proxies for BBs were associated with lower HDL-cholesterol and higher triacylglycerols. For CCBs, the genetic proxies were related to higher LDL-cholesterol in both ancestries.
In a further assessment of their roles in lipid fractions and lipoprotein fractions, genetic proxies for BBs were also related to lower HDL-cholesterol, and consistently associated with lower concentrations of HDL particles (Fig. 3). In line with the association of BBs with higher triacylglycerols (Fig. 3), genetic proxies for BBs were also related to higher triacylglycerol fractions (VLDL-triacylglycerols and LDL-triacylglycerols). In addition, there were positive associations with VLDL-cholesterol and VLDL particle concentrations. For genetically predicted ACE inhibitor and CCBs, the associations with lipid fractions, lipoprotein fractions or apolipoproteins were not statistically significant after Bonferroni correction (Fig. 3). The result of the power calculation is shown in ESM Table 5. At 80% power and α = 0.05, a smaller effect size was detected in Europeans than in East Asians.
In sensitivity analysis using another set of genetic predictors derived from the UK Biobank, we observed similar estimates (ESM Figs 3 and 4), although the CI for BBs and diabetes included the null (ESM Fig. 3), and there was a positive association of BBs with LDL-cholesterol (ESM Fig. 4). The estimates were also robust to the use of different analysis methods (a weighted median and MR-PRESSO) (ESM Tables 6 and 7). Steiger filtering indicated directionality from genetic predictors of antihypertensive drugs to diabetes, glucose and lipids. In the assessment of the metabolic effect of SBP, we found that genetically predicted SBP was related to higher risk of diabetes and higher triacylglycerols in Europeans (ESM Table 8) and had null associations in East Asians (ESM Table 9).
Systematic review and meta-analysis
There were 1327 studies in the initial search. After screening, we identified six trials comparing ACE inhibitors vs placebo or CCBs, and three trials comparing BBs vs placebo or CCBs (ESM Table 10). There is large heterogeneity between studies (I2 of 71.6% for the former and 71.3% for the latter). Using a random-effects model, individuals taking ACE inhibitors had a lower risk of diabetes (ESM Fig. 5a), and those taking BBs had a higher risk of diabetes (ESM Fig. 5b). The funnel plot indicated no publication bias for ACE inhibitors (ESM Fig. 6a), but possibly two missing studies for BBs (ESM Fig. 6b). Sensitivity analysis using trim and fill showed consistent findings (risk ratio 1.42, 95% CI 1.18–1.70). All studies were of high quality (ESM Table 11), which suggests that any heterogeneity is not due to differences in study quality.
Discussion
Using MR to minimise residual confounding, this study suggests that genetically predicted ACE inhibition is associated with a lower risk of diabetes in Europeans and East Asians. Echoing the previous concerns about BBs [39, 40], this MR study shows that genetically predicted BBs may relate to a higher risk of diabetes and an unfavourable lipid profile. Moreover, genetic proxies for CCBs are related to higher LDL-cholesterol in both Europeans and East Asians, but the clinical significance of the small effect size is unclear.
This trans-ancestry MR study shows different associations of different classes of antihypertensive drugs with diabetes, blood glucose levels and lipid profile. The pleiotropic associations of genetically proxied ACE inhibition and BBs with diabetes in Europeans were supported by a phenome-wide scan of pleiotropic effects based on the UK Biobank [16]. In comparison with the previous scan of disease outcomes in Europeans [16], the current study focused on the metabolic off-target effects, with in-depth evaluation of the effect of antihypertensive drugs on both diabetes and metabolic traits. This study also integrated GWAS in Europeans and East Asians, and extended the evidence in Europeans to East Asians. The pleiotropic associations of ACE inhibitors and BBs with diabetes in this MR were corroborated by our meta-analysis of RCTs, although the RCTs for BBs and diabetes mainly report side-effects or secondary outcomes. In comparison with RCTs, which are often performed in people with hyperglycaemia [4], MR estimates in generally healthy participants extend the evidence by showing that the benefits of ACE inhibitors on diabetes and blood glucose levels may also apply to the general population.
Despite the protective role of these antihypertensive drugs, BBs and CCBs have shown an unfavourable association with glucose metabolism and/or lipid profile in the current study. Consistent with small trials using BBs [41], the associations with HDL-cholesterol and triacylglycerols for BBs in our MR study provides more evidence supporting long-term concerns regarding BBs [42]. Notably, the patterns of associations in East Asians suggest that the concerns raised in Europeans also apply to East Asians. In the further assessment of lipid fractions and lipoprotein subclasses, we found similar changes in HDL- and triacylglycerol-related fractions and particle concentrations. These associations confirm the unfavourable effect of BBs on HDL-cholesterol and triacylglycerols. Moreover, genetic proxies for CCBs may link to higher LDL-cholesterol in Europeans and East Asians. However, the associations of CCBs with lipid fractions and lipoprotein subclasses were generally null, which requires replication.
Given their common effect on lowering BP, the different associations for the various classes of antihypertensive drugs suggest that mechanisms beyond lowering BP exist. In our study, genetically predicted SBP was related to higher risk of diabetes and higher triacylglycerols in Europeans, indicating that the antihypertensive effect of the drugs is expected to lower diabetes and triacylglycerols. As such, the association of genetically predicted SBP cannot explain the observed unfavourable effects of BBs and CCBs on diabetes or lipids. Currently, the mechanisms underlying the varying metabolic associations have not been clarified. The positive association of genetically proxied CCBs with LDL-cholesterol is in accordance with a previous interventional study [10], but the mechanism is obscure. BBs may affect energy expenditure [43] and lower insulin sensitivity [7], which may explain or partly explain the unfavourable metabolic effect. BBs also lower testosterone in animal experiments [44]. An MR study has suggested that genetically predicted higher level of testosterone is related to a lower risk of diabetes in men [45], which may partly explain the association with diabetes for BBs, but this pathway remains to be examined in humans. ACE inhibitors may also play a role by modulating sex hormones. Specifically, ACE inhibitors increase sex hormone-binding globulin levels in women [46], and accumulating evidence shows sex hormone-binding globulin may lower the risk of diabetes [45].
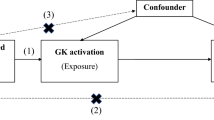
Despite its novelty, this study has several limitations. First, MR relies on three assumptions: that the genetic instruments are associated with the exposure, are not related to potential confounders, and the association of the genetic instruments with the outcome is exclusively through affecting the exposure [47]. To satisfy these assumptions, we used genetic instruments in drug target genes related to SBP in Europeans and East Asians. The consistent patterns of association using the two sets of genetic proxies in Europeans and East Asians provide more confidence to the findings. Second, weak instruments may bias towards the null; however, we only used genetic variants with F-statistic >10. Third, MR estimates, although less confounded, are less precise than conventional observational studies, because the genetic variants only explain a small proportion of the variance in exposure [33]. As such, we cannot exclude the possibility that the null associations are due to a lack of precision, especially in East Asians. Replication in larger studies of East Asians would be worthwhile. Fourth, for the analysis with glucose in UK Biobank and Biobank Japan, the genetic associations with proxies for antihypertensive drugs and with glucose were obtained from the same study, which may bias the estimates [48]. However, we used by far the largest GWAS in this study, and the validity of overlapping samples in large cohorts was supported in a recent simulation study [49]. Fifth, genetic effects may be diluted by compensatory processes or feedback mechanisms, which may mitigate genetic effects and bias the estimates towards the null [50]. However, the directions of associations should be unchanged. The directions of associations were also corroborated by systematic review and meta-analyses, although most trials included did not use diabetes as the primary outcome. Sixth, MR studies assess lifelong associations, and the magnitude of effect may not be comparable to the short-term effects of taking antihypertensive drugs, hence this study is more relevant in assessing the direction of associations than determining the magnitude of associations.
From the perspective of clinical practice, our findings suggest that, compared with BBs and CCBs, ACE inhibitors, which have beneficial association with diabetes and neutral association with lipids, are more suitable for the management of hypertension with complications of the metabolic syndrome. In the era of evidence-based medicine, our findings from a large trans-ancestry MR provide another piece of evidence complementary to clinical trials, as references for hypertension management. Given the wide prevalence of hypertension and the lack of large RCTs in East Asians, these findings provided timely evidence regarding healthcare of hypertension. Moreover, a deeper understanding of the mechanisms underlying the off-target metabolic associations would be worthwhile, providing insights for new drug development and repositioning.
Conclusions
Our findings suggest protective association of genetically predicted ACE inhibition with diabetes and glycaemic traits, while genetic proxies for BBs may be related to higher risk of diabetes. Genetic proxies for BBs and CCBs may also be related to an unfavourable lipid profile in Europeans and East Asians. Understanding the mechanisms underlying these diverse associations of different classes of antihypertensive drugs would be valuable, with implications for drug development and repositioning, as well as developing optimal strategies for the prevention and treatment of hypertension in people with diabetes and/or hyperlipidaemia.
Data availability
All data sources used in this study are included in the ESM and are also available from the corresponding authors on reasonable request. This systematic review and meta-analysis was registered in Research Registry (number: reviewregistry1239), providing details of the literature search, selection criteria, data extraction and data analysis.
Abbreviations
- BB:
-
β-Blocker
- CAD:
-
Coronary artery disease
- CCB:
-
Calcium channel blocker
- GWAS:
-
Genome-wide association studies
- LD:
-
Linkage disequilibrium
- MR:
-
Mendelian randomisation
- MR-PRESSO:
-
Mendelian Randomisation Pleiotropy RESidual Sum and Outlier
- SBP:
-
Systolic blood pressure
References
Hansson L, Lindholm LH, Niskanen L et al (1999) Effect of angiotensin-converting-enzyme inhibition compared with conventional therapy on cardiovascular morbidity and mortality in hypertension: the captopril prevention project (CAPPP) randomised trial. Lancet 353(9153):611–616. https://doi.org/10.1016/s0140-6736(98)05012-0
Blackburn DF, Wilson TW (2006) Antihypertensive medications and blood sugar: theories and implications. Can J Cardiol 22(3):229–233. https://doi.org/10.1016/s0828-282x(06)70902-3
NICE (2019) NICE guideline [NG136]. Available from www.nice.org.uk/guidance/ng136/chapter/Recommendations#choosing-antihypertensive-drug-treatment-for-people-with-or-without-type-2-diabetes
DREAM Trial Investigators, Bosch J, Yusuf S et al (2006) Effect of ramipril on the incidence of diabetes. N Engl J Med 355(15):1551–1562. https://doi.org/10.1056/NEJMoa065061
Liou YS, Chen HY, Tien L, Gu YS, Jong GP (2015) Antihypertensive drug use and new-onset diabetes in female patients with coronary artery disease: a population-based longitudinal cohort study. Medicine (Baltimore) 94(36):e1495. https://doi.org/10.1097/MD.0000000000001495
Gress TW, Nieto FJ, Shahar E, Wofford MR, Brancati FL (2000) Hypertension and antihypertensive therapy as risk factors for type 2 diabetes mellitus. Atherosclerosis risk in communities study. N Engl J Med 342(13):905–912. https://doi.org/10.1056/NEJM200003303421301
Lee WG, Murphy R, McCall JL et al (2017) Nadolol reduces insulin sensitivity in liver cirrhosis: a randomized double-blind crossover trial. Diabetes Metab Res Rev 33(3). https://doi.org/10.1002/dmrr.2859
Racine N, Hamet P, Sampalis JS, Longo N, Bastien N (2010) A 52-week prospective, cohort study of the effects of losartan with or without hydrochlorothiazide (HCTZ) in hypertensive patients with metabolic syndrome. J Hum Hypertens 24(11):739–748. https://doi.org/10.1038/jhh.2010.3
Black HR, Davis B, Barzilay J et al (2008) Metabolic and clinical outcomes in nondiabetic individuals with the metabolic syndrome assigned to chlorthalidone, amlodipine, or lisinopril as initial treatment for hypertension: a report from the antihypertensive and lipid-lowering treatment to prevent heart attack trial (ALLHAT). Diabetes Care 31(2):353–360. https://doi.org/10.2337/dc07-1452
Rachmani R, Lidar M, Brosh D, Levi Z, Ravid M (2000) Oxidation of low-density lipoprotein in normotensive type 2 diabetic patients. Comparative effects of enalapril versus nifedipine: a randomized cross-over over study. Diabetes Res Clin Pract 48(2):139–145. https://doi.org/10.1016/s0168-8227(99)00149-7
Woo KS, Norris RM, Nicholls G (1995) Racial difference in incidence of cough with angiotensin-converting enzyme inhibitors (a tale of two cities). Am J Cardiol 75(14):967–968. https://doi.org/10.1016/s0002-9149(99)80703-6
Ma RC, Chan JC (2013) Type 2 diabetes in east Asians: similarities and differences with populations in Europe and the United States. Ann N Y Acad Sci 1281:64–91. https://doi.org/10.1111/nyas.12098
Ference BA (2018) How to use Mendelian randomization to anticipate the results of randomized trials. Eur Heart J 39(5):360–362. https://doi.org/10.1093/eurheartj/ehx462
Gill D, Georgakis MK, Walker VM et al (2021) Mendelian randomization for studying the effects of perturbing drug targets. Wellcome Open Res 6:16. https://doi.org/10.12688/wellcomeopenres.16544.2
Levin MG, Klarin D, Walker VM et al (2021) Association between genetic variation in blood pressure and increased lifetime risk of peripheral artery disease. Arterioscler Thromb Vasc Biol 41(6):2027–2034. https://doi.org/10.1161/ATVBAHA.120.315482
Gill D, Georgakis MK, Koskeridis F et al (2019) Use of genetic variants related to antihypertensive drugs to inform on efficacy and side effects. Circulation 140(4):270–279. https://doi.org/10.1161/CIRCULATIONAHA.118.038814
Georgakis MK, Gill D, Webb AJS et al (2020) Genetically determined blood pressure, antihypertensive drug classes, and risk of stroke subtypes. Neurology 95(4):e353–e361. https://doi.org/10.1212/WNL.0000000000009814
Zhao JV, Schooling CM (2021) Using Mendelian randomization study to assess the renal effects of antihypertensive drugs. BMC Med 19(1):79. https://doi.org/10.1186/s12916-021-01951-4
Pigeyre M, Sjaarda J, Chong M et al (2020) ACE and type 2 diabetes risk: a Mendelian randomization study. Diabetes Care 43(4):835–842. https://doi.org/10.2337/dc19-1973
Aschard H, Vilhjalmsson BJ, Joshi AD, Price AL, Kraft P (2015) Adjusting for heritable covariates can bias effect estimates in genome-wide association studies. Am J Hum Genet 96(2):329–339. https://doi.org/10.1016/j.ajhg.2014.12.021
Kanai M, Akiyama M, Takahashi A et al (2018) Genetic analysis of quantitative traits in the Japanese population links cell types to complex human diseases. Nat Genet 50(3):390–400. https://doi.org/10.1038/s41588-018-0047-6
Lu X, Wang L, Lin X et al (2015) Genome-wide association study in Chinese identifies novel loci for blood pressure and hypertension. Hum Mol Genet 24(3):865–874. https://doi.org/10.1093/hmg/ddu478
Takeuchi F, Akiyama M, Matoba N et al (2018) Interethnic analyses of blood pressure loci in populations of east Asian and European descent. Nat Commun 9(1):5052. https://doi.org/10.1038/s41467-018-07345-0
Mahajan A, Taliun D, Thurner M et al (2018) Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat Genet 50(11):1505–1513. https://doi.org/10.1038/s41588-018-0241-6
Spracklen CN, Horikoshi M, Kim YJ et al (2020) Identification of type 2 diabetes loci in 433,540 east Asian individuals. Nature 582(7811):240–245. https://doi.org/10.1038/s41586-020-2263-3
Sudlow C, Gallacher J, Allen N et al (2015) UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med 12(3):e1001779. https://doi.org/10.1371/journal.pmed.1001779
Willer CJ, Schmidt EM, Sengupta S et al (2013) Discovery and refinement of loci associated with lipid levels. Nat Genet 45(11):1274–1283. https://doi.org/10.1038/ng.2797
UK Biobank (2018) Nightingale Health and UK Biobank announces major initiative to analyse half a million blood samples to facilitate global medical research. Available from www.ukbiobank.ac.uk/learn-more-about-uk-biobank/news/nightingale-health-and-uk-biobank-announces-major-initiative-to-analyse-half-a-million-blood-samples-to-facilitate-global-medical-research
Lu X, Huang J, Mo Z et al (2016) Genetic susceptibility to lipid levels and lipid change over time and risk of incident hyperlipidemia in Chinese populations. Circ Cardiovasc Genet 9(1):37–44. https://doi.org/10.1161/CIRCGENETICS.115.001096
van der Harst P, Verweij N (2018) Identification of 64 novel genetic loci provides an expanded view on the genetic architecture of coronary artery disease. Circ Res 122(3):433–443. https://doi.org/10.1161/CIRCRESAHA.117.312086
Ishigaki K, Akiyama M, Kanai M et al (2020) Large-scale genome-wide association study in a Japanese population identifies novel susceptibility loci across different diseases. Nat Genet 52(7):669–679. https://doi.org/10.1038/s41588-020-0640-3
Lu X, Peloso GM, Liu DJ et al (2017) Exome chip meta-analysis identifies novel loci and east Asian-specific coding variants that contribute to lipid levels and coronary artery disease. Nat Genet 49(12):1722–1730. https://doi.org/10.1038/ng.3978
Burgess S (2014) Sample size and power calculations in Mendelian randomization with a single instrumental variable and a binary outcome. Int J Epidemiol 43(3):922–929. https://doi.org/10.1093/ije/dyu005
Freeman G, Cowling BJ, Schooling CM (2013) Power and sample size calculations for Mendelian randomization studies using one genetic instrument. Int J Epidemiol 42(4):1157–1163. https://doi.org/10.1093/ije/dyt110
Hemani G, Tilling K, Smith GD (2017) Orienting the causal relationship between imprecisely measured traits using GWAS summary data. PLoS Genet 13(11):e1007081. https://doi.org/10.1371/journal.pgen.1007081
Bowden J, Davey Smith G, Haycock PC, Burgess S (2016) Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol 40(4):304–314. https://doi.org/10.1002/gepi.21965
Verbanck M, Chen CY, Neale B, Do R (2018) Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet 50(5):693–698. https://doi.org/10.1038/s41588-018-0099-7
Verhagen AP, de Vet HC, de Bie RA et al (1998) The Delphi list: a criteria list for quality assessment of randomized clinical trials for conducting systematic reviews developed by Delphi consensus. J Clin Epidemiol 51(12):1235–1241. https://doi.org/10.1016/S0895-4356(98)00131-0
Bangalore S, Parkar S, Grossman E, Messerli FH (2007) A meta-analysis of 94,492 patients with hypertension treated with beta blockers to determine the risk of new-onset diabetes mellitus. Am J Cardiol 100(8):1254–1262. https://doi.org/10.1016/j.amjcard.2007.05.057
Belknap S (2008) Review: β-blockers for hypertension increase risk of new onset diabetes. Evid Based Med 13(2):50. https://doi.org/10.1136/ebm.13.2.50
Boquist S, Ruotolo G, Hellenius ML, Danell-Toverud K, Karpe F, Hamsten A (1998) Effects of a cardioselective β-blocker on postprandial triglyceride-rich lipoproteins, low density lipoprotein particle size and glucose-insulin homeostasis in middle-aged men with modestly increased cardiovascular risk. Atherosclerosis 137(2):391–400. https://doi.org/10.1016/s0021-9150(97)00278-5
Moser M (1989) Relative efficacy of, and some adverse reactions to, different antihypertensive regimens. Am J Cardiol 63(4):2B–7B. https://doi.org/10.1016/0002-9149(89)90931-4
Lee P, Kengne AP, Greenfield JR, Day RO, Chalmers J, Ho KK (2011) Metabolic sequelae of β-blocker therapy: weighing in on the obesity epidemic? Int J Obes 35(11):1395–1403. https://doi.org/10.1038/ijo.2010.284
Segarra AB, Prieto I, Villarejo AB et al (2013) Effects of antihypertensive drugs on angiotensinase activities in the testis of spontaneously hypertensive rats. Horm Metab Res 45(5):344–348. https://doi.org/10.1055/s-0032-1329988
Ruth KS, Day FR, Tyrrell J et al (2020) Using human genetics to understand the disease impacts of testosterone in men and women. Nat Med 26(2):252–258. https://doi.org/10.1038/s41591-020-0751-5
Koshida H, Takeda R, Miyamori I (1998) Lisinopril decreases plasma free testosterone in male hypertensive patients and increases sex hormone binding globulin in female hypertensive patients. Hypertens Res 21(4):279–282. https://doi.org/10.1291/hypres.21.279
Davies NM, Holmes MV, Davey Smith G (2018) Reading Mendelian randomisation studies: a guide, glossary, and checklist for clinicians. BMJ 362:k601. https://doi.org/10.1136/bmj.k601
Burgess S, Davies NM, Thompson SG (2016) Bias due to participant overlap in two-sample Mendelian randomization. Genet Epidemiol 40(7):597–608. https://doi.org/10.1002/gepi.21998
Minelli C, Del Greco MF, van der Plaat DA, Bowden J, Sheehan NA, Thompson J (2021) The use of two-sample methods for Mendelian randomization analyses on single large datasets. Int J Epidemiol 50(5):1651–1659. https://doi.org/10.1093/ije/dyab084
Lawlor DA, Harbord RM, Sterne JAC, Timpson N, Davey-Smith G (2008) Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat Med 27(8):1133–1163. https://doi.org/10.1002/Sim.3034
Acknowledgements
The authors would like to thank all studies and consortia (listed in ESM Table 1) for providing the valuable data.
Authors’ relationships and activities
The authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work.
Funding
This work was supported by the National Natural Science Foundation of China (91857118, 81773537, 82030102) and the Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (2021-I2M-1-010, 2019-I2M-2-003).
Author information
Authors and Affiliations
Contributions
JVZ conceived the original idea; JVZ and XFL designed the study. JVZ analysed the data and interpreted the findings with the help of XFL, FCL, CMS, JXL and DFG. JVZ drafted the manuscript; XFL, FCL, CMS, JXL and DFG revised the manuscript. All authors gave final approval of the manuscript. JVZ and XFL are the guarantors of this study.
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 793 kb)
Rights and permissions
About this article
Cite this article
Zhao, J.V., Liu, F., Schooling, C.M. et al. Using genetics to assess the association of commonly used antihypertensive drugs with diabetes, glycaemic traits and lipids: a trans-ancestry Mendelian randomisation study. Diabetologia 65, 695–704 (2022). https://doi.org/10.1007/s00125-021-05645-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-021-05645-7